Protein Model Discrimination Using Mutational Sensitivity Derived from Deep Sequencing - ScienceDirect
3D Model Structure of Predicted Protein Illustrated by I-TASSER Software | Download Scientific Diagram
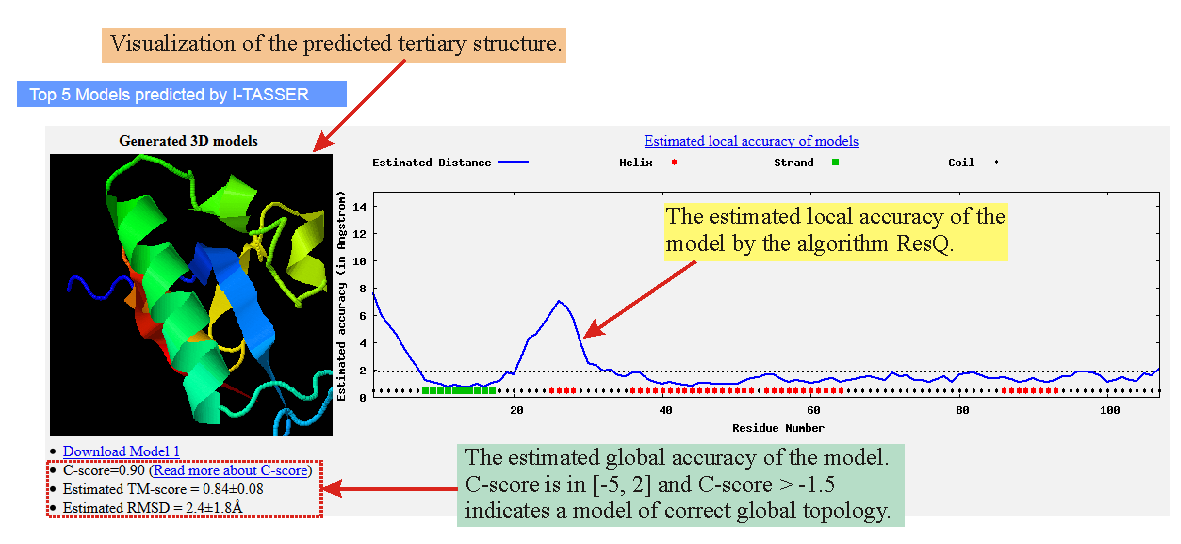
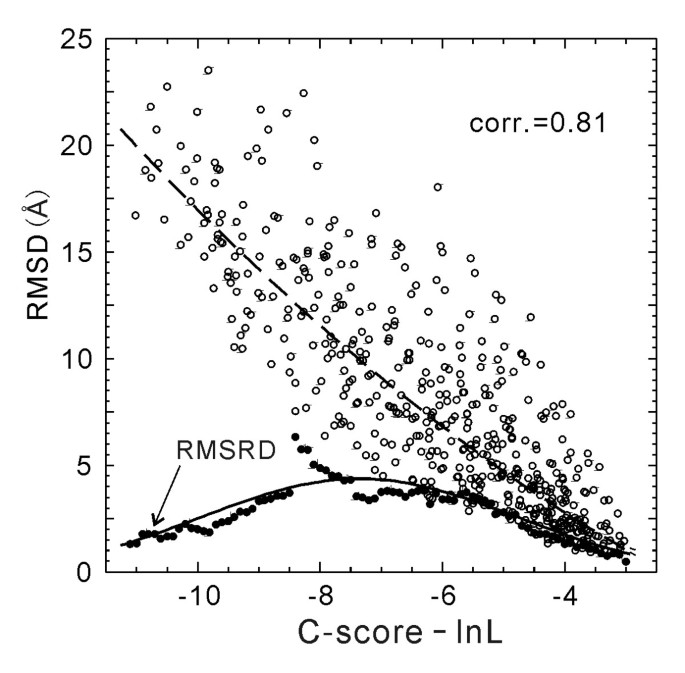
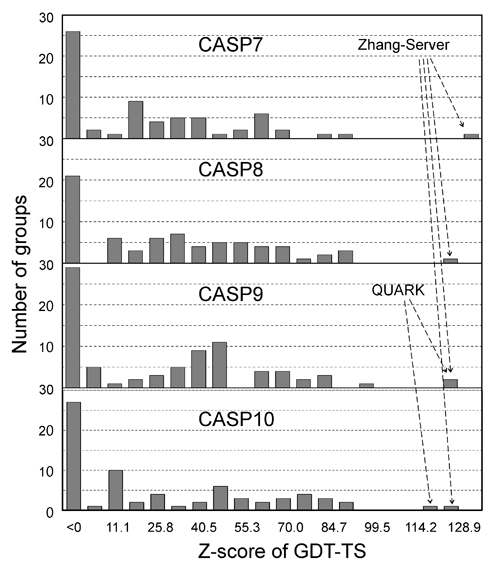
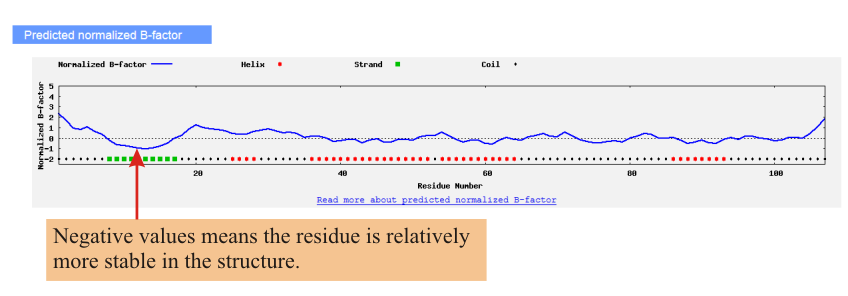
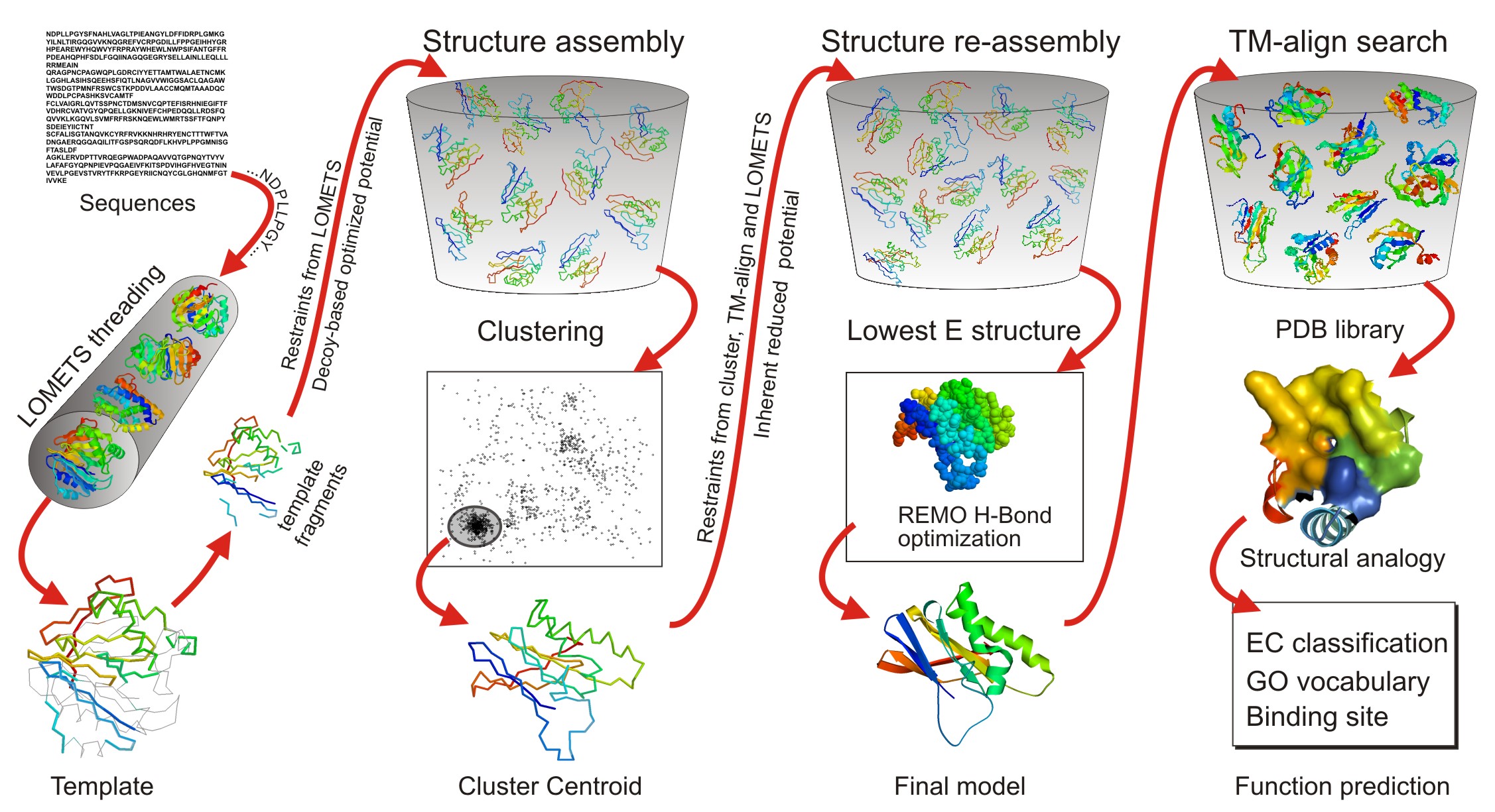
![PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/682fc6a2dcf6edaadacb1a2f90fdd137aef28d15/2-Figure1-1.png)
PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar

β-Barrels covalently link peptidoglycan and the outer membrane in the α-proteobacterium Brucella abortus | Nature Microbiology

I-TASSER gateway: A protein structure and function prediction server powered by XSEDE. - Abstract - Europe PMC

Protein structure, amino acid composition and sequence determine proteome vulnerability to oxidation‐induced damage | The EMBO Journal

Improving Protein Structure Prediction Using Multiple Sequence-Based Contact Predictions - ScienceDirect

I-TASSER gateway: A protein structure and function prediction server powered by XSEDE - ScienceDirect

I-TASSER gateway: A protein structure and function prediction server powered by XSEDE. - Abstract - Europe PMC